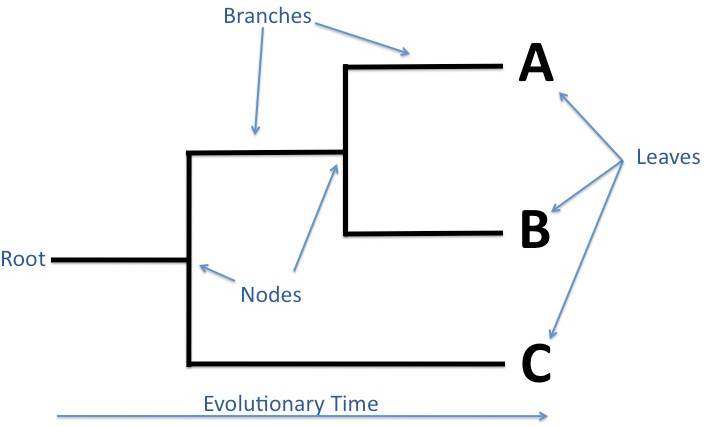
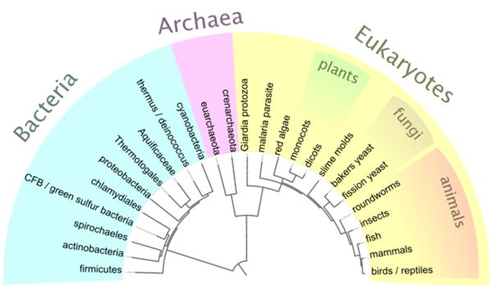
What is phylogeny?
The number of mutations between two organisms can tell us how closely related the two species are. The study of these mutations can also give us an indication of the time that has elapsed since the species' diverged. Comparative biology and the study of homologous traits among species, is relevant in helping to distinguish the different level of relatedness among all organisms [1]. From all of this information, phylogenies and phylogenetic trees may be created to help visualize the evolution of these genetically related group of organisms.
What is AMPK-gamma-2's protein phylogeny?
Phylogenetic trees were created using the AMPK-gamma-2 protein homologs [as seen on the Protein Homology Page]. These phylogenetic trees were created using ClustalOmega percent identity and BLOSUM62 matrices that help determine the path of evolution with the fewest evolutionary events. Each program uses their own specific algorithm to account for possible insertions and deletions that may happen throughout evolution.
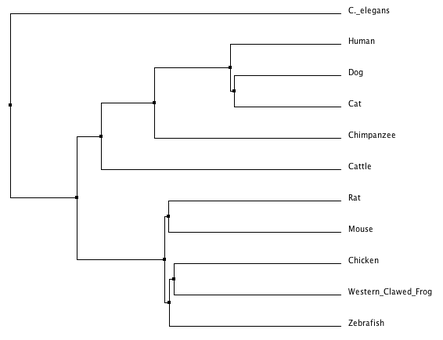
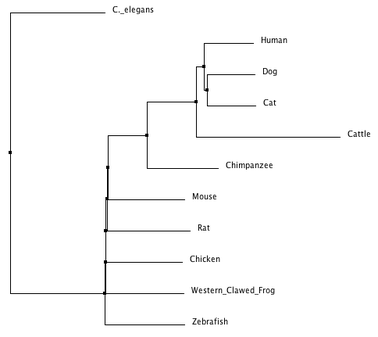
There are two different kind of phylogenetic trees that can be created. An average distance tree uses the specific amino acid sequences in order to create a tree that has the least amount of dissimilarity between its' cluster members, while the neighbor joining tree method produces an unrooted tree that does not require the assumption of a constant rate of evolution [2].
Phylogenetic trees can also be constructed using either a BLOSUM62 or % Identity matrix. The BLOSUM62 matrix uses substitution scores based on the chance that one amino acid would be exchanged for another, while the % Identity matrix uses the actual DNA sequences to determine the amount of similarity between the homologs.
There are two different kind of phylogenetic trees that can be created. An average distance tree uses the specific amino acid sequences in order to create a tree that has the least amount of dissimilarity between its' cluster members, while the neighbor joining tree method produces an unrooted tree that does not require the assumption of a constant rate of evolution [2].
Phylogenetic trees can also be constructed using either a BLOSUM62 or % Identity matrix. The BLOSUM62 matrix uses substitution scores based on the chance that one amino acid would be exchanged for another, while the % Identity matrix uses the actual DNA sequences to determine the amount of similarity between the homologs.
Analysis
Despite being produced using the same database (ClustalOmega), there are some discrepancies between the two phylogenetic trees that were created. Both trees parallel in their alignment of the human AMPK-gamma-2 protein being closest in relationship to it's homolog in the cat and dog. Both trees also place the C. elegans homologous protein as having the amino acid sequence most dissimilar from the rest of the organisms. The most obvious discrepancy is where each tree placed the cattle and the chimpanzee. The average distance tree placed the chimpanzee homolog more closely related to the human than the cattle homolog, yet the neighbor joining tree has cattle closer than chimpanzee. This discrepancy suggests that there was some sort of amino acid change in the cattle and chimpanzee protein that each method evaluated in a very different manner. BLOSUM62 seemed to be the appropriate matrix to use when analyzing the protein phylogeny because it is based on the likelihood of amino acid substitutions and it's different scores reflect estimates of what constitutes conserved substitutions in the evolution of these proteins [3].
References
[1] n.d. Plant and Animal Evolution. Retrieved from the University of Waikato School of Science and Engineering Website: http://sci.waikato.ac.nz/evolution/Homology.shtml
[2] Opperdoes, Fred. "The Neighbor-Joining Method." de Duve Institute of Cellular Pathology, ICP. 08 Aug. 1997. Web. 10 May 2013. <http://www.icp.ucl.ac.be/~opperd/private/neighbor.html>
[3] http://www.ebi.ac.uk/help/matrix.html#BLOSSUM
[2] Opperdoes, Fred. "The Neighbor-Joining Method." de Duve Institute of Cellular Pathology, ICP. 08 Aug. 1997. Web. 10 May 2013. <http://www.icp.ucl.ac.be/~opperd/private/neighbor.html>
[3] http://www.ebi.ac.uk/help/matrix.html#BLOSSUM
Margaret Beatka ([email protected])
Page Last Updated: 5/10/13
This web page was produced as an assignment for Genetics 677, as an undergraduate course at UW-Madison.
Page Last Updated: 5/10/13
This web page was produced as an assignment for Genetics 677, as an undergraduate course at UW-Madison.