What is DNA homology?
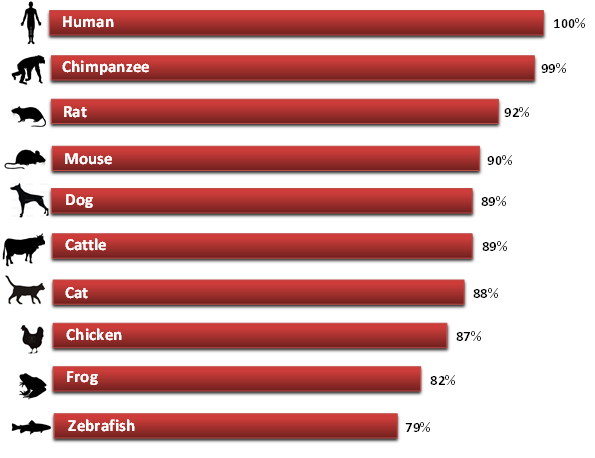
The study of DNA homology is an examination of the relatedness of certain species that is attributable to a common origin. Genes may also be orthologous to one another if throughout speciation they have retained function. Two species that share similar DNA sequences most often have a common ancestor and common evolutionary ancestry [1]. Understanding and determining gene homologs is very important in conducting human disease research due to the fact that it is unethical to experiment on humans. Comparative biology such as this enables researchers to determine the most appropriate model organism that contains the most similar homolog to the human gene of interest.
What are the homologs of the human prkag2 gene?
|
Putative homologs for the human prkag2 gene were found using the program BLAST. BLAST is a program that interprets sequences of genomic data and compares it to other species with sequenced genomes. It displays the results of other known species with significant DNA alignments and also allows you to pair up your sequence with any other species you're interested in [2]. The file below contains the mRNA sequences (in FASTA format) of the human prkag2 gene compared to the homologous mRNA sequences of thirteen other organisms (six of them being common research model organisms).
|
|||||||
List of Homologous Genes
|
Homo sapiens (Humans) - PRKAG2
Accession Number: AJ249976.1 GI: 6688198 FASTA Pan troglodytes (Chimpanzee) - PRKAG2 Accession Number: XM_003318924.1 GI: 332870103 E Value: 0.0 Max. Identity: 99% FASTA Rattus norvegicus (Rat) -Prkag2 Accession Number: AY376854.1 GI: 38604627 Max. Identity: 92% E Value: 0.0 FASTA Mus musculus (Mouse) - Prkag2 Accession Number: NM_001170555.1 GI: 282847326 E Value: 0.0 Max. Identity: 90% FASTA Canis lupus familiaris (Dog) - PRKAG2 Accession Number: XM_532769.3 GI: 345781414 E Value: 0.0 Max. Identity: 89% FASTA Bos taurus (Cattle) - PRKAG2 Accession Number: XM_002686979.2 GI: 359065132 E Value: 0.0 Max. Identity: 89% FASTA Felis catus (Cat) - PRKAG2 Accession Number: XM_003983233.1 GI: 410953241 Max. Identity: 88% FASTA Gallus gallus (Chicken) - PRKAG2 Accession Number: DQ212711.1 GI: 77158184 E Value: 0.0 Max. Identity: 87% FASTA Xenopus tropicalis (Clawed Frog) - PRKAG1 Accession Number: NM_001127939.1 GI: 189230173 E Value: 0.0 Max. Identity: 82% FASTA Dan rerio (Zebrafish) - zgc:153329 Accession Number: NM_001077179.1 GI: 116004574 E Value: 0.0 Max. Identity: 79% FASTA Caenorhabditis elegans - aakg-1 Accession Number: NM_067236.4 GI: 392896977 FASTA |
What are the alignments of PRKAG2 gene?
Multiple sequence alignments were performed on all 11 organisms using ClustalW2 and T-Coffee. These programs align multiple DNA sequences to help identify conserved regions and help determine evolutionary relationships between organisms.
|
|
||||||||||||
Analysis
The prkag2 gene seems to be very well conserved among many of the other vertebrates it was BLASTed against, such as the chimpanzee, mouse, and rat. This can be determined from the very small E value's that were produced when the human prkag2 gene was put through BLAST. These small E values suggest a more significant score and alignment because they represent the number of different alignments with scores equivalent to or better than what is expected to occur by chance [2]. In regards to invertebrates, Zebrafish returned a result that suggested this prkag2 gene may also be somewhat conserved in these organisms too. Although WormBase identified the C.elegans gene (aakg-1) to be orthologous to the human PRKAG2 gene, BLAST was unable to identify any "highly similar sequences" among the two. The fact that this gene appears to be highly conserved among many organisms suggests that this gene is unique and essential [3].
References
[1] n.d. Plant and Animal Evolution. Retrieved from the University of Waikato School of Science and Engineering Website: http://sci.waikato.ac.nz/evolution/Homology.shtml
[2] Altschul S.F., Gish W., Miller W., Myers E.W. and Lipman D.J. (1990) Basic local alignment search tool. Journal of Molecular Biology. 215: 403-410.
[3] http://www.wormbase.org/db/get?name=WBGene00013732;class=Gene
[2] Altschul S.F., Gish W., Miller W., Myers E.W. and Lipman D.J. (1990) Basic local alignment search tool. Journal of Molecular Biology. 215: 403-410.
[3] http://www.wormbase.org/db/get?name=WBGene00013732;class=Gene
Margaret Beatka ([email protected])
Page Last Updated: 5/10/13
This web page was produced as an assignment for Genetics 677, as an undergraduate course at UW-Madison.
Page Last Updated: 5/10/13
This web page was produced as an assignment for Genetics 677, as an undergraduate course at UW-Madison.