What is gene ontology?
Gene ontology is the process of developing a structured and consistent vocabulary the describes gene products in three different categories: biological processes, cellular components, and molecular functions. It is an effort to provide specific annotations of orthologous genes across human and model organism genomes [1]. Biological processes ontology categorizes a specific series of events, within a living organism, that has a defined beginning and end. The molecular functions ontology refers to the specific abilities and functions of a gene product that occur at the molecular level. The last category, cellular component ontology, is used to describe the locations and levels of subcellular structures and macromolecular complexes of a gene product [2].
Gene ontologies of specific gene products can be browsed, visualized, and researched using web applications such as AmiGO.
Gene ontologies of specific gene products can be browsed, visualized, and researched using web applications such as AmiGO.
What are the gene ontologies of prkag2?
|
Biological Processes
ATP biosynthetic process, carnitine shuttle, cell cycle arrest, cellular lipid metabolic process, energy reserve metabolic process, fatty acid biosynthetic process, glycogen metabolic process, insulin receptor signaling pathway, intracellular protein kinase cascade, negative regulation of protein kinase activity, positive regulation of peptidyl-threonine phosphorylation, positive regulation of protein kinase activity, regulation of fatty acid oxidation, regulation of glucose import, regulation of glycolysis, small molecular metabolic process, and sterol biosynthetic process. |
Molecular Functions
ADP binding, ATP binding, cAMP-dependent protein kinase inhibitor activity, cAMP-dependent protein kinase regulator activity, phosphorylase kinase regulator activity, protein kinase activator activity, and protein kinase binding. Cellular Components
AMP-activated protein kinase complex, cytosol, and nucleoplasm. |
PRKAG2 Gene Ontology Flowcharts
|
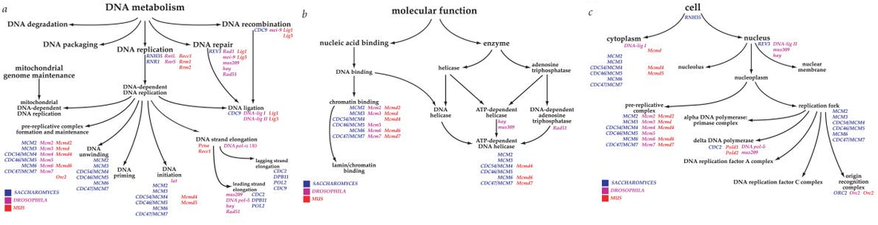
The three gene ontology diagrams located to the right and below were obtained from QuickGo, a gene ontology browser that presents flowcharts showing how a specific biological process, molecular function, or cellular component interacts with other processes, pathways, and/or components of the cell.
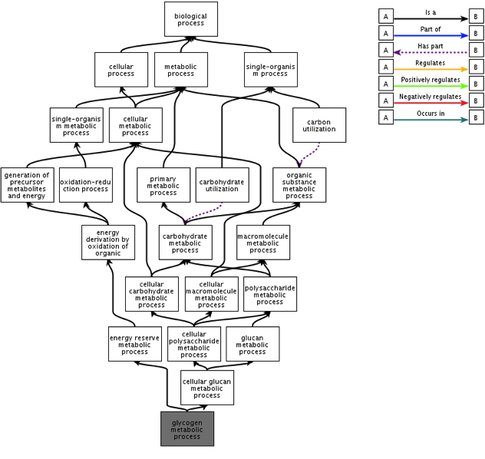
Glycogen metabolism (Figure 2) is a biological process that is important to illustrate for the prkag2 gene and it's resulting protein due to the belief that this process may give rise to supraventricular tachycardia--a prevailing symptom of WPW syndrome [3]. The complexity of the glycogen metabolism process can be seen quite obviously due to the large number of processes, and extensive connections between these processes that are portrayed in the flowchart. This suggests that the gene and protein are involved in a substantial and variable number of biological processes. |
|
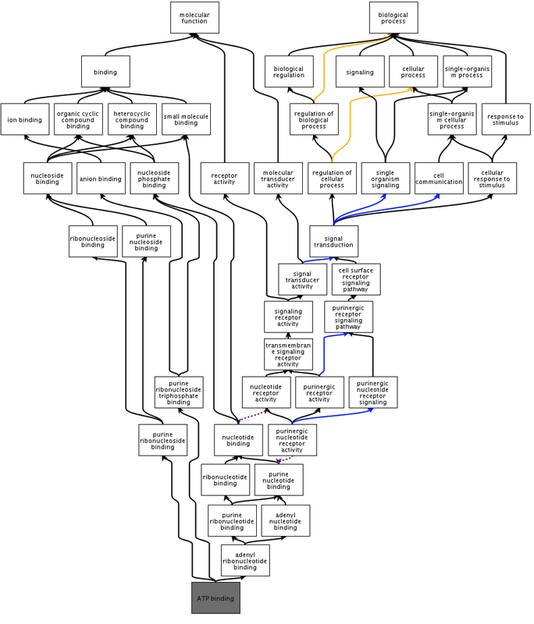
ATP binding is a molecular function that is universally important due to its involvement in the selective interactions of ATP, an extremely important enzyme regulator and coenzyme. ATP binding may be impaired in certain prkag2 gene mutations and cause WPW symptoms. Figure 3 shows how the prkag2 protein is involved in ATP binding as a molecular function and biological process.
|
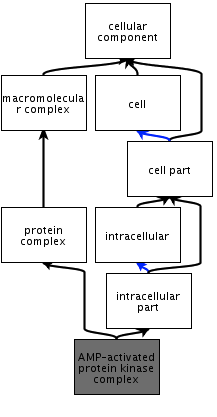
The AMP-activated protein kinase complex is a protein complex that plays an enzymatic role in maintaining the cellular energy homeostasis of the cell. The gamma subunit of this AMPK complex is the specific subunit that is affected when there are mutations to the prkag2 gene. Figure 4 shows the cellular components of this AMPK complex as a whole.
Analysis
The complexity of the glycogen metabolism and ATP binding gene ontology flowcharts suggest that prkag2 is a highly conserved gene with important biological functions that have been extensively studied and is well understood. The resulting gene ontology terms reiterate what we already know about the function of prkag2 and that is involved in processes regarding glycogen and ATP, among others.
References
[1] The Gene Ontology Consortium. (2008). The gene ontology project in 2008. Nucleic Acids Research, 36(1):D440-4. doi: 10.1093/nar/gkm883
[2] Ashburner, M., Ball, C. A., Blake, J. A., et al. (2000). Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genetics, 25: 25-9. doi: 10.1038/75556.
[3] Burwinkel, B., Scott, J.W., Buhrer, C., et al. (2005). Fatal congentital heart glycogenosis caused by a recurrent activating R531Q mutation in the gamma2-subunit of AMP-activated protein kinase (PRKAG2), not by phosphorylase kinase deficiency. The American Journal of Human Genetics. 76(6): 1034-49. doi: 10.1086/430840.
[4] Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S, AmiGO Hub, Web Presence Working Group. AmiGO: online access to ontology and annotation data. Bioinformatics. Jan 2009;25(2):288-9. doi:
[2] Ashburner, M., Ball, C. A., Blake, J. A., et al. (2000). Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genetics, 25: 25-9. doi: 10.1038/75556.
[3] Burwinkel, B., Scott, J.W., Buhrer, C., et al. (2005). Fatal congentital heart glycogenosis caused by a recurrent activating R531Q mutation in the gamma2-subunit of AMP-activated protein kinase (PRKAG2), not by phosphorylase kinase deficiency. The American Journal of Human Genetics. 76(6): 1034-49. doi: 10.1086/430840.
[4] Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S, AmiGO Hub, Web Presence Working Group. AmiGO: online access to ontology and annotation data. Bioinformatics. Jan 2009;25(2):288-9. doi:
Margaret Beatka ([email protected])
Page Last Updated: 4/1/13
This web page was produced as an assignment for Genetics 677, as an undergraduate course at UW-Madison.
Page Last Updated: 4/1/13
This web page was produced as an assignment for Genetics 677, as an undergraduate course at UW-Madison.